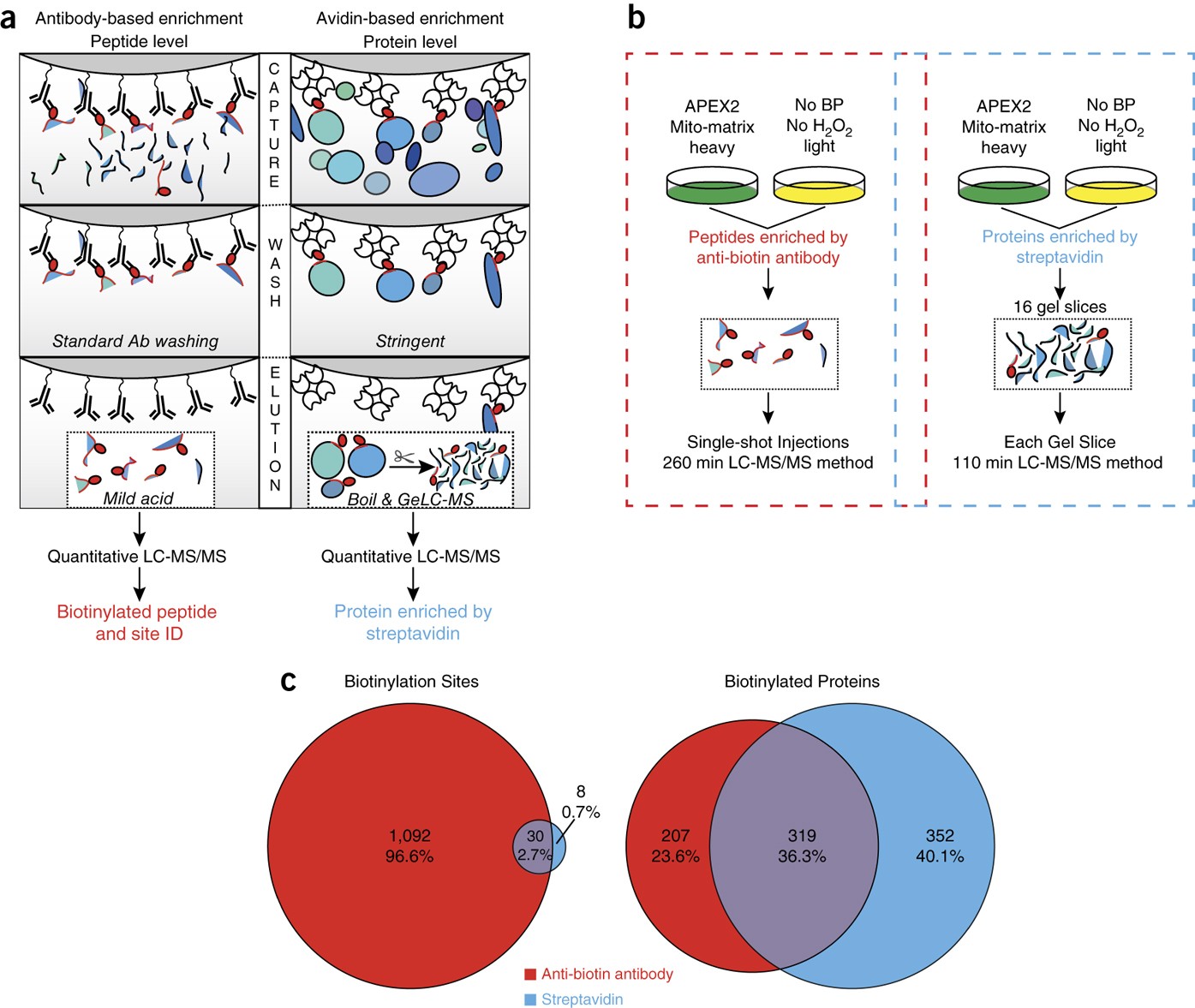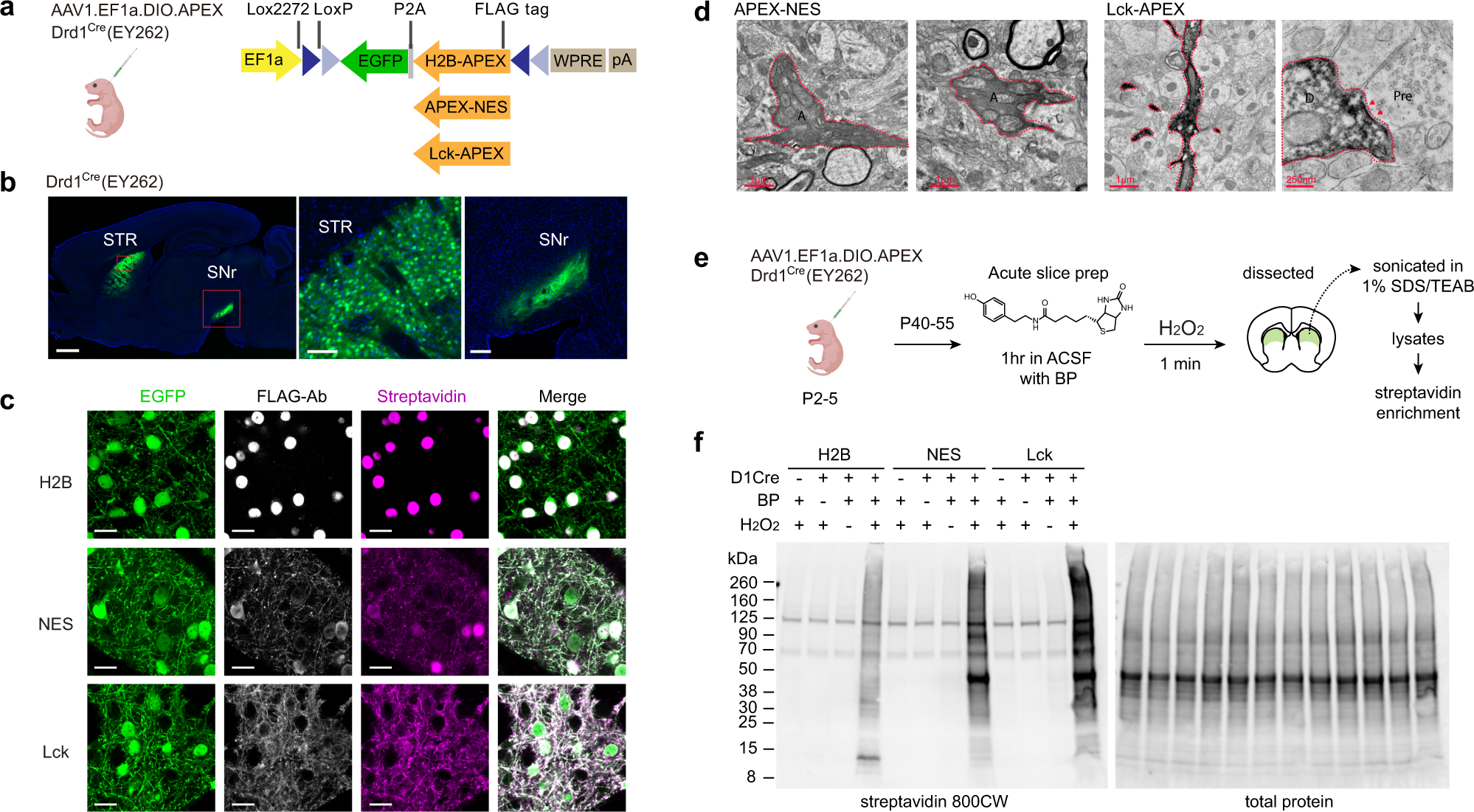
Cell-type and subcellular compartment-specific APEX2 proximity labeling reveals activity-dependent nuclear proteome dynamics in the striatum | Nature Communications
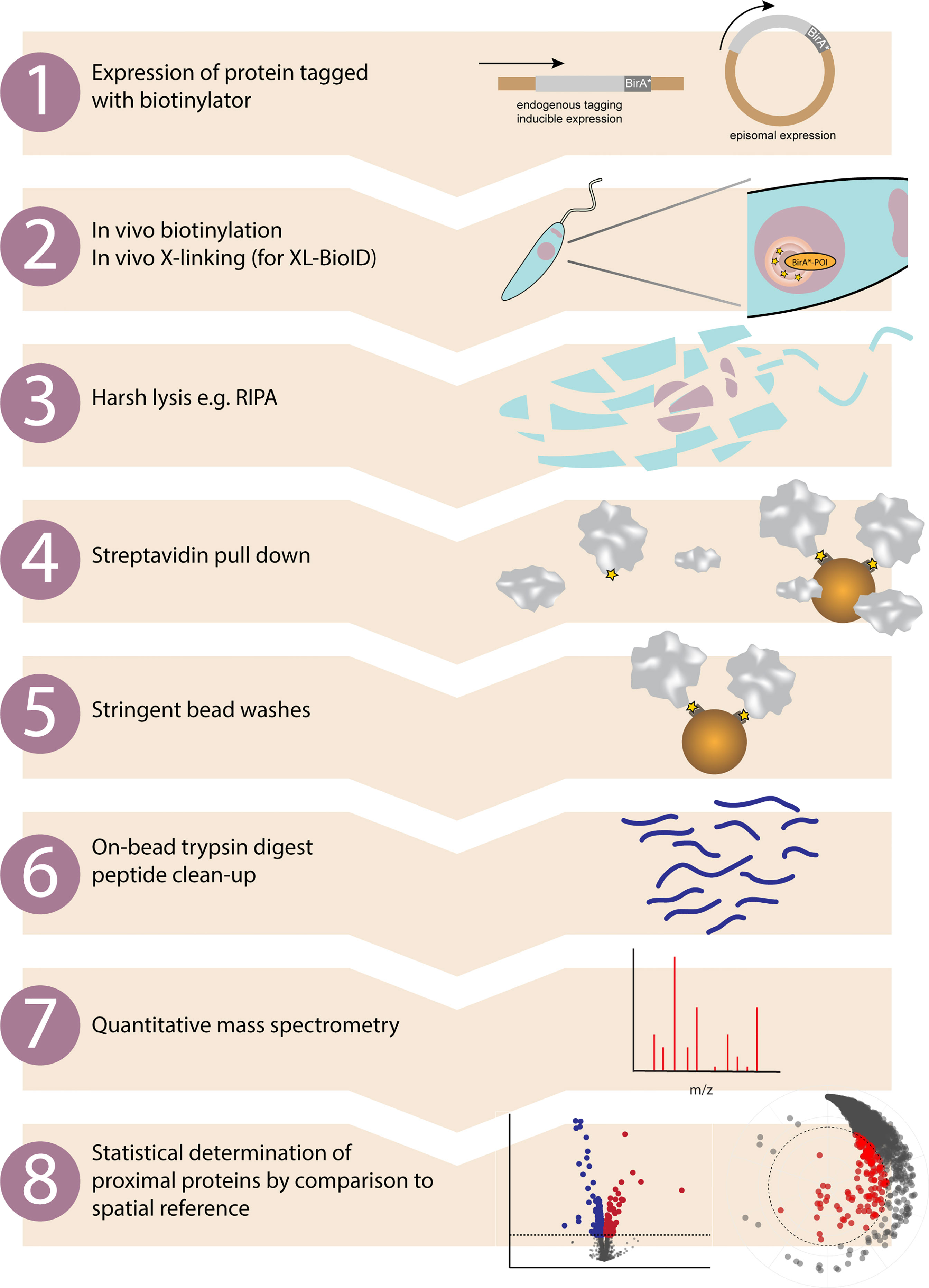
Frontiers | Tag Thy Neighbour: Nanometre-Scale Insights Into Kinetoplastid Parasites With Proximity Dependent Biotinylation

Avidin–Biotin Technology in Gold Nanoparticle-Decorated Graphene Field Effect Transistors for Detection of Biotinylated Macromolecules with Ultrahigh Sensitivity and Specificity | ACS Omega

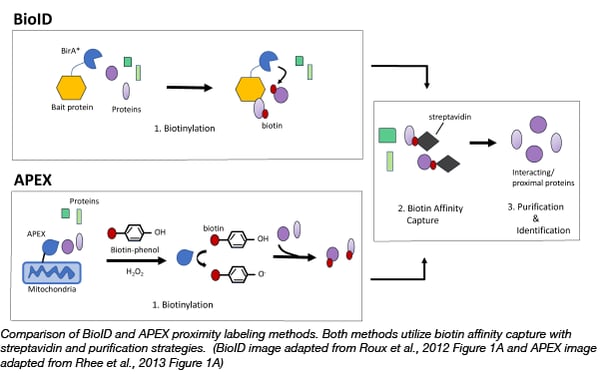
Meet the neighbors: Mapping local protein interactomes by proximity‐dependent labeling with BioID - Varnaitė - 2016 - PROTEOMICS - Wiley Online Library

Proximity Biotinylation as a Method for Mapping Proteins Associated with mtDNA in Living Cells - ScienceDirect

Proximity‐dependent biotinylation approaches to study apicomplexan biology - Kimmel - 2022 - Molecular Microbiology - Wiley Online Library
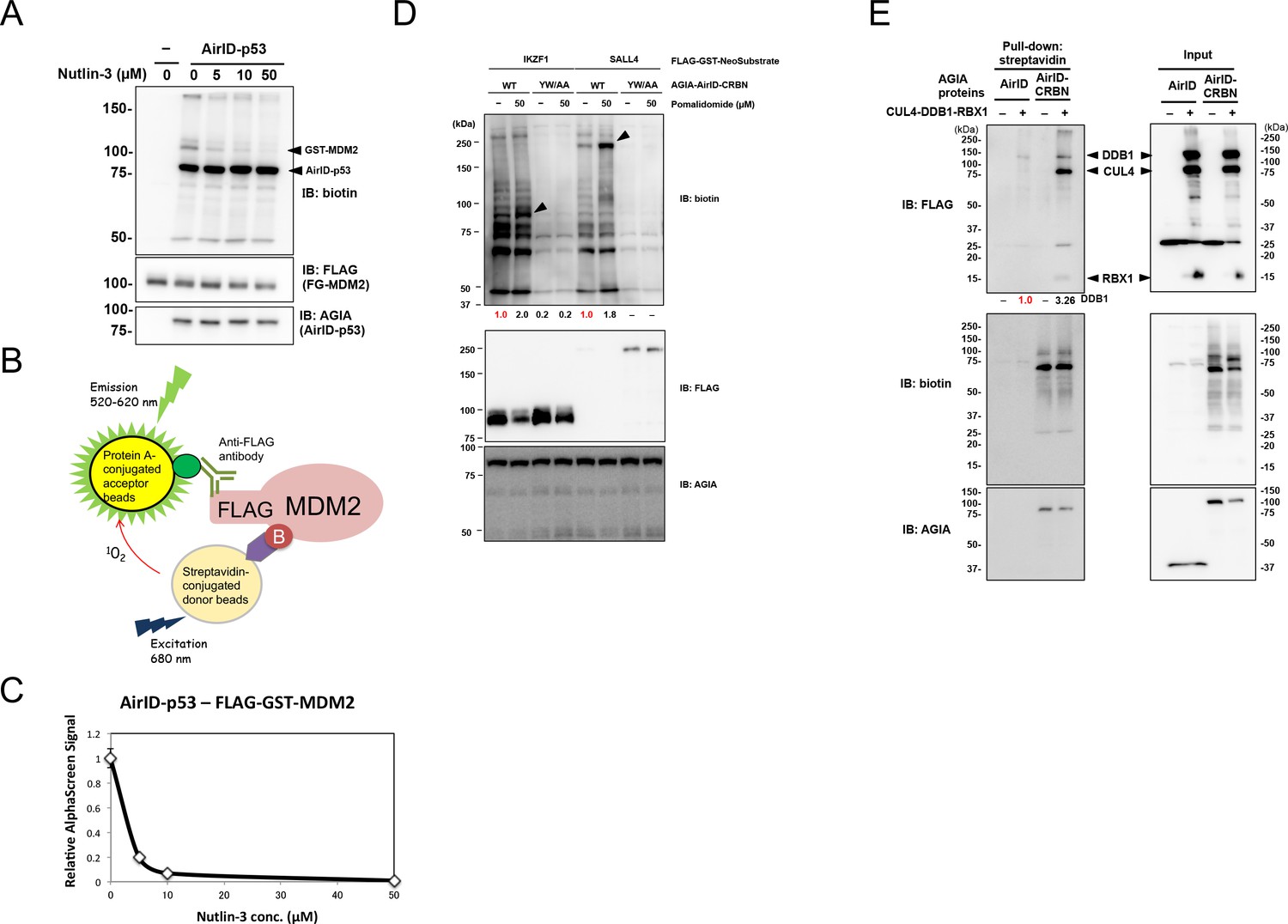
AirID, a novel proximity biotinylation enzyme, for analysis of protein– protein interactions | eLife

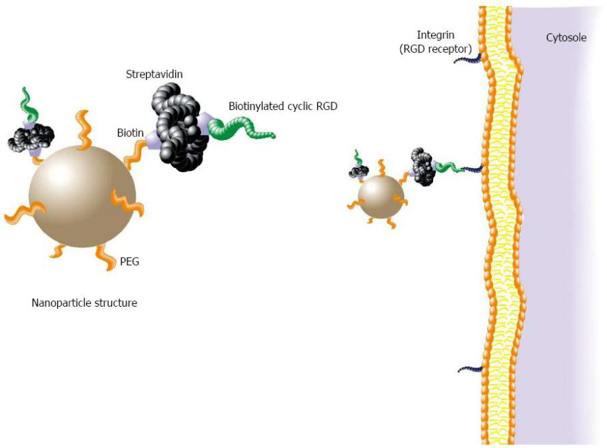
Targeting of biotinylated compounds to its target tissue using a low-density lipoprotein receptor–avidin fusion protein | Gene Therapy

Electron Microscopic Detection of Single Membrane Proteins by a Specific Chemical Labeling - ScienceDirect

Streptavidin staining of biotinylated proteins reveals proteins at the... | Download Scientific Diagram

Structure-Based Design with Tag-Based Purification and In-Process Biotinylation Enable Streamlined Development of SARS-CoV-2 Spike Molecular Probes - ScienceDirect

Optimized protocol for the identification of lipid droplet proteomes using proximity labeling proteomics in cultured human cells: STAR Protocols

Comparison of the biotin detection system with EM autoradiography and... | Download Scientific Diagram

MicroID2: A Novel Biotin Ligase Enables Rapid Proximity-Dependent Proteomics - Molecular & Cellular Proteomics