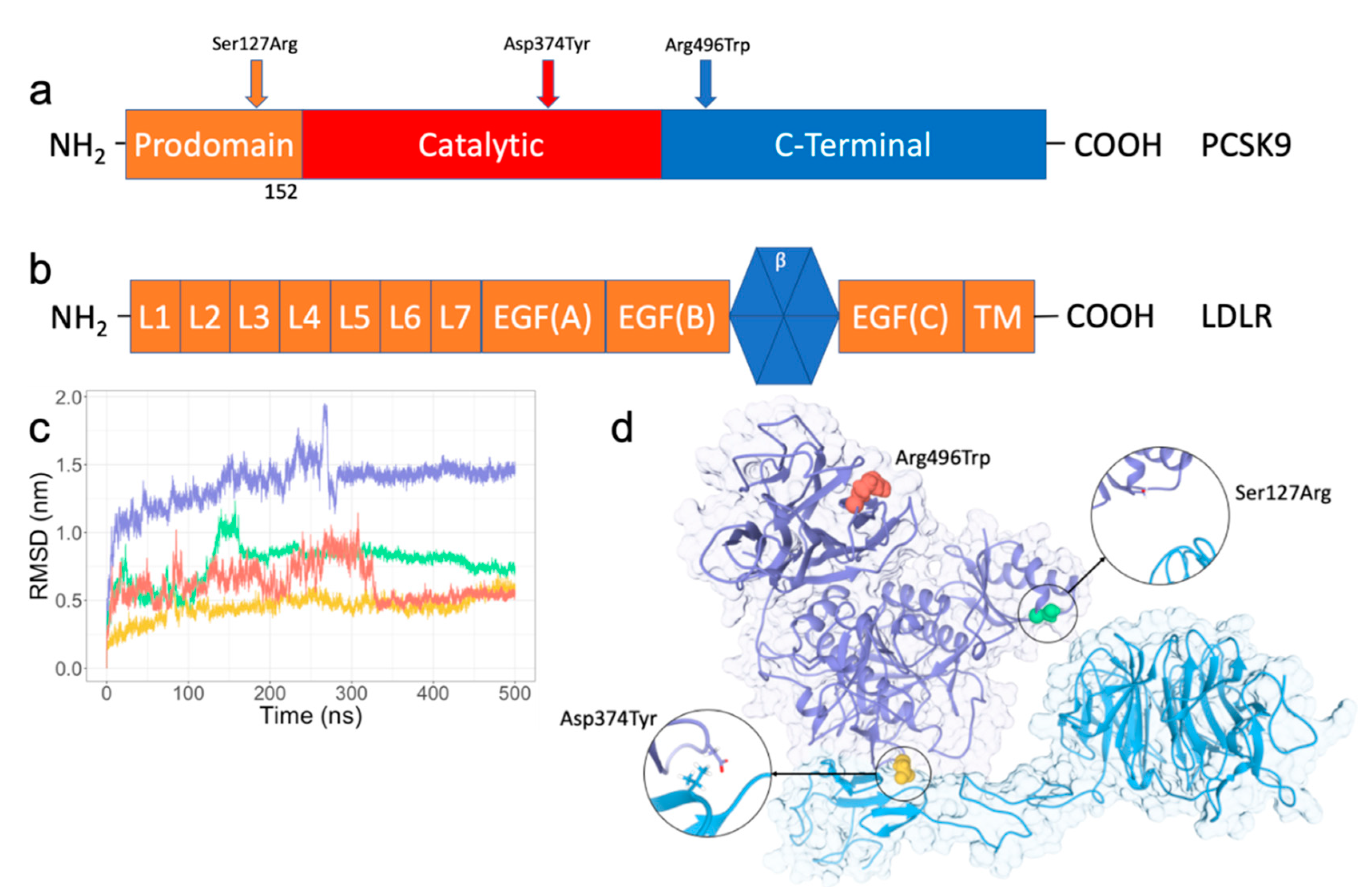
IJMS | Free Full-Text | In Silico Insights into Protein–Protein Interaction Disruptive Mutations in the PCSK9-LDLR Complex | HTML
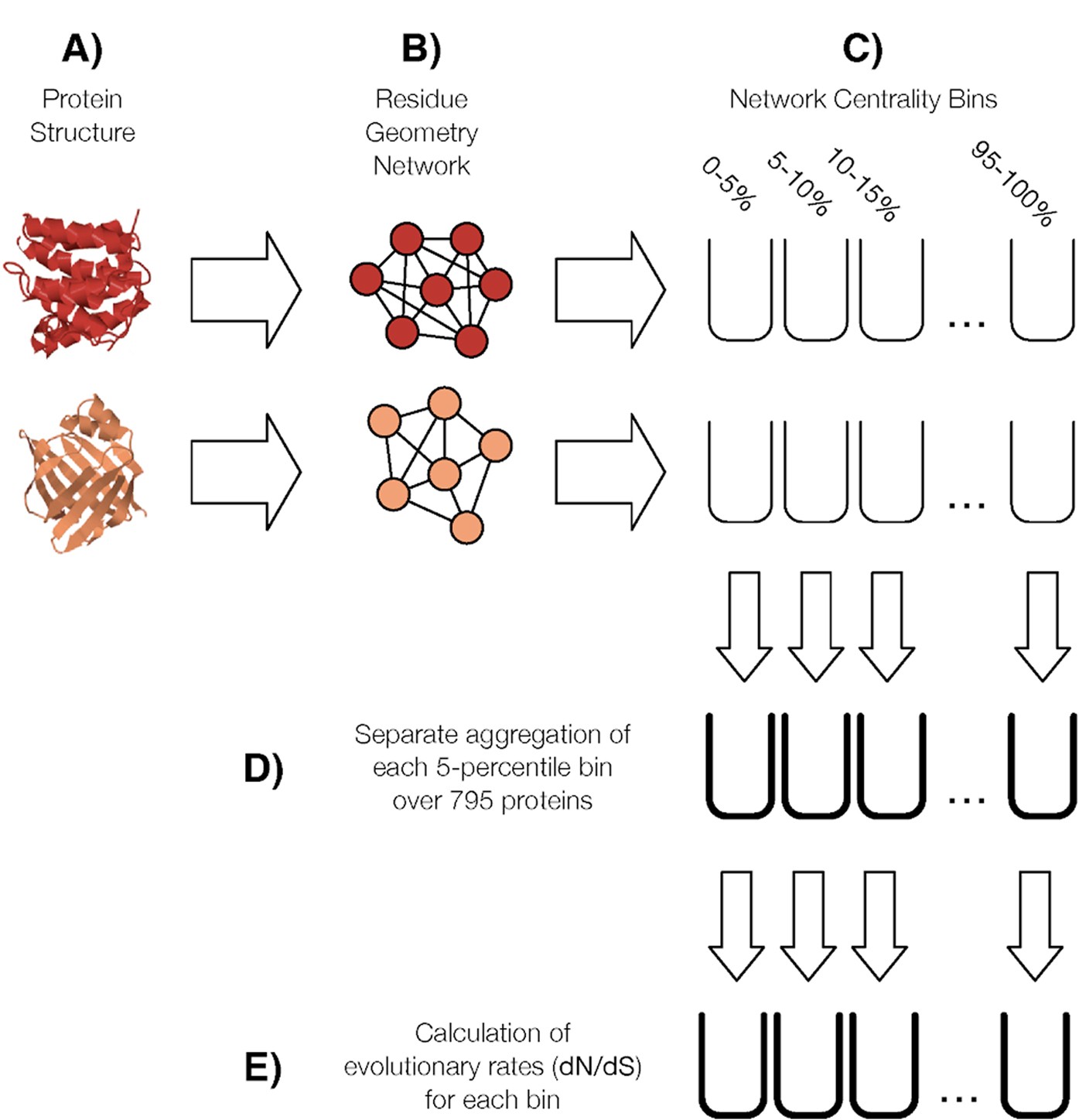
Residue Geometry Networks: A Rigidity-Based Approach to the Amino Acid Network and Evolutionary Rate Analysis | Scientific Reports

AlphaFold2 models indicate that protein sequence determines both structure and dynamics | Scientific Reports

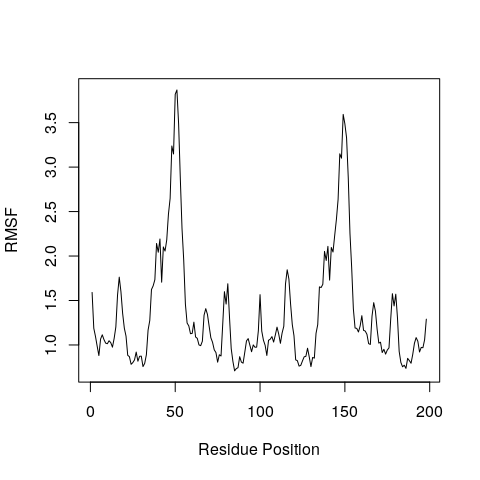
Example for how to calculate the root mean square fluctuation (RMSF)... | Download Scientific Diagram

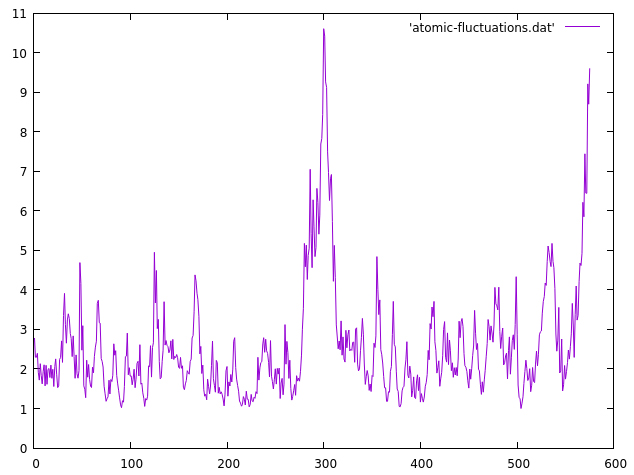
Figure S3. Per-residue fluctuation of the b-factor during simulation... | Download Scientific Diagram
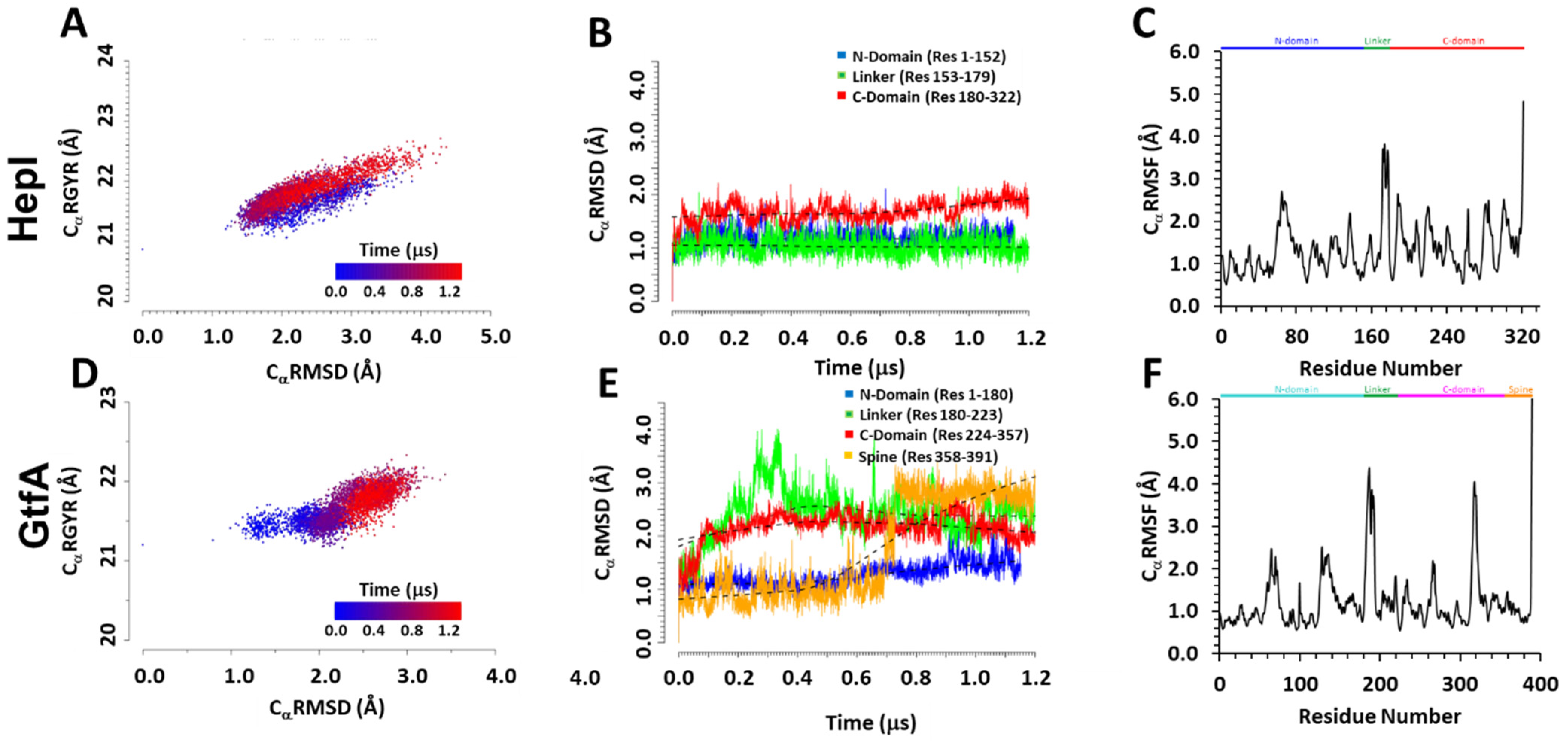
IJMS | Free Full-Text | Conserved Conformational Hierarchy across Functionally Divergent Glycosyltransferases of the GT-B Structural Superfamily as Determined from Microsecond Molecular Dynamics | HTML
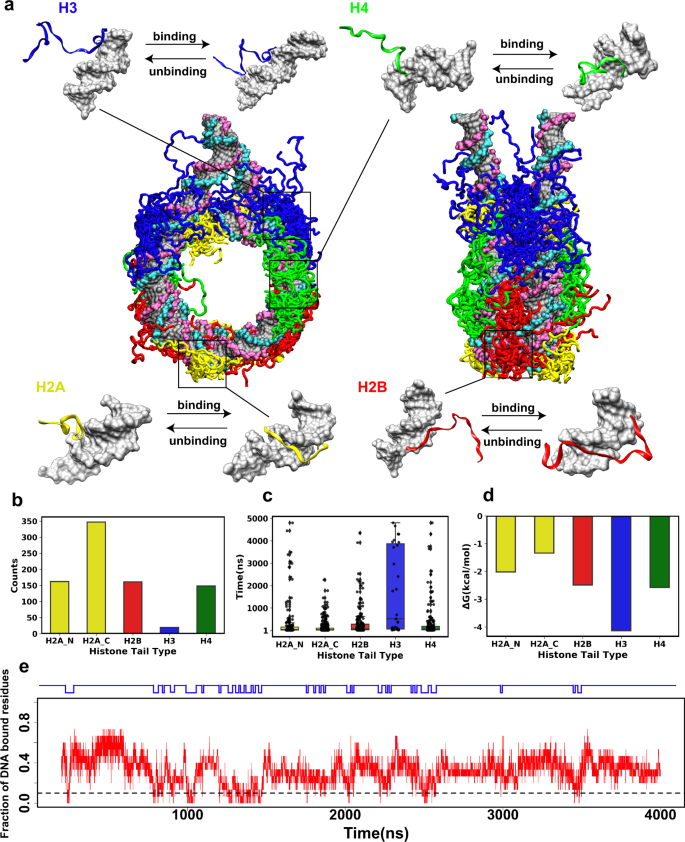
Binding of regulatory proteins to nucleosomes is modulated by dynamic histone tails | Nature Communications

Enhanced Conformational Sampling with an Adaptive Coarse-Grained Elastic Network Model Using Short-Time All-Atom Molecular Dynamics | Journal of Chemical Theory and Computation

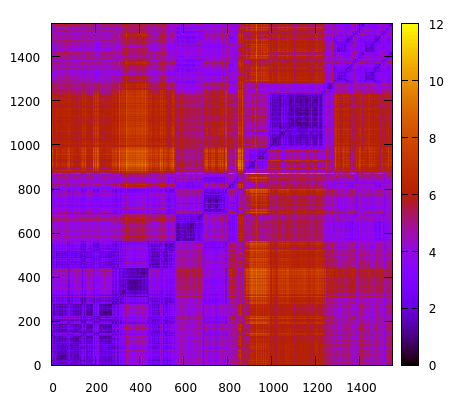
Comparison of residue fluctuation (root mean square fluctuation (RMSF))... | Download Scientific Diagram

Root Mean Square Fluctuation. Per-residue root mean square fluctuation... | Download Scientific Diagram
Docking studies and molecular dynamics simulations of the binding characteristics of waldiomycin and its methyl ester analog to Staphylococcus aureus histidine kinase | PLOS ONE
Computational drug repurposing strategy predicted peptide-based drugs that can potentially inhibit the interaction of SARS-CoV-2 spike protein with its target (humanACE2) | PLOS ONE
![Dynamic residue interaction network analysis of the oseltamivir binding site of N1 neuraminidase and its H274Y mutation site conferring drug resistance in influenza A virus [PeerJ] Dynamic residue interaction network analysis of the oseltamivir binding site of N1 neuraminidase and its H274Y mutation site conferring drug resistance in influenza A virus [PeerJ]](https://peerj.com/articles/11552/graphical-abstract.jpg)
Dynamic residue interaction network analysis of the oseltamivir binding site of N1 neuraminidase and its H274Y mutation site conferring drug resistance in influenza A virus [PeerJ]

Secondary Structure and Conformational Stability of the Antigen Residues Making Contact with Antibodies | The Journal of Physical Chemistry B
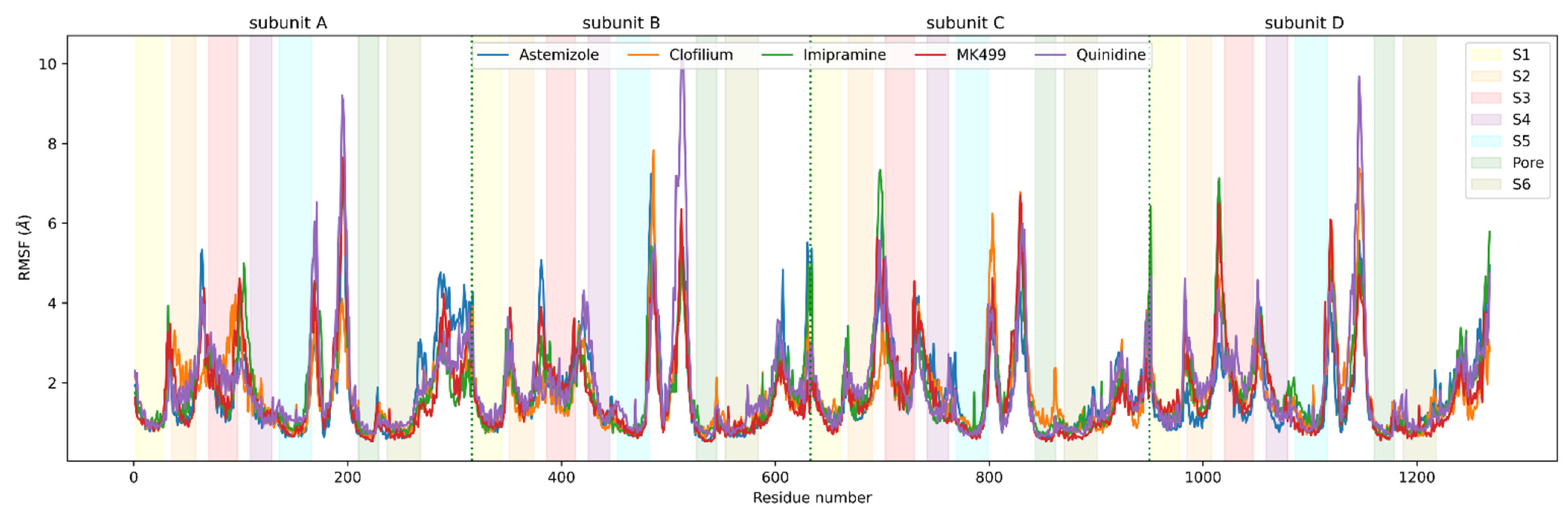
IJMS | Free Full-Text | Molecular Dynamics-Derived Pharmacophore Model Explaining the Nonselective Aspect of KV10.1 Pore Blockers | HTML
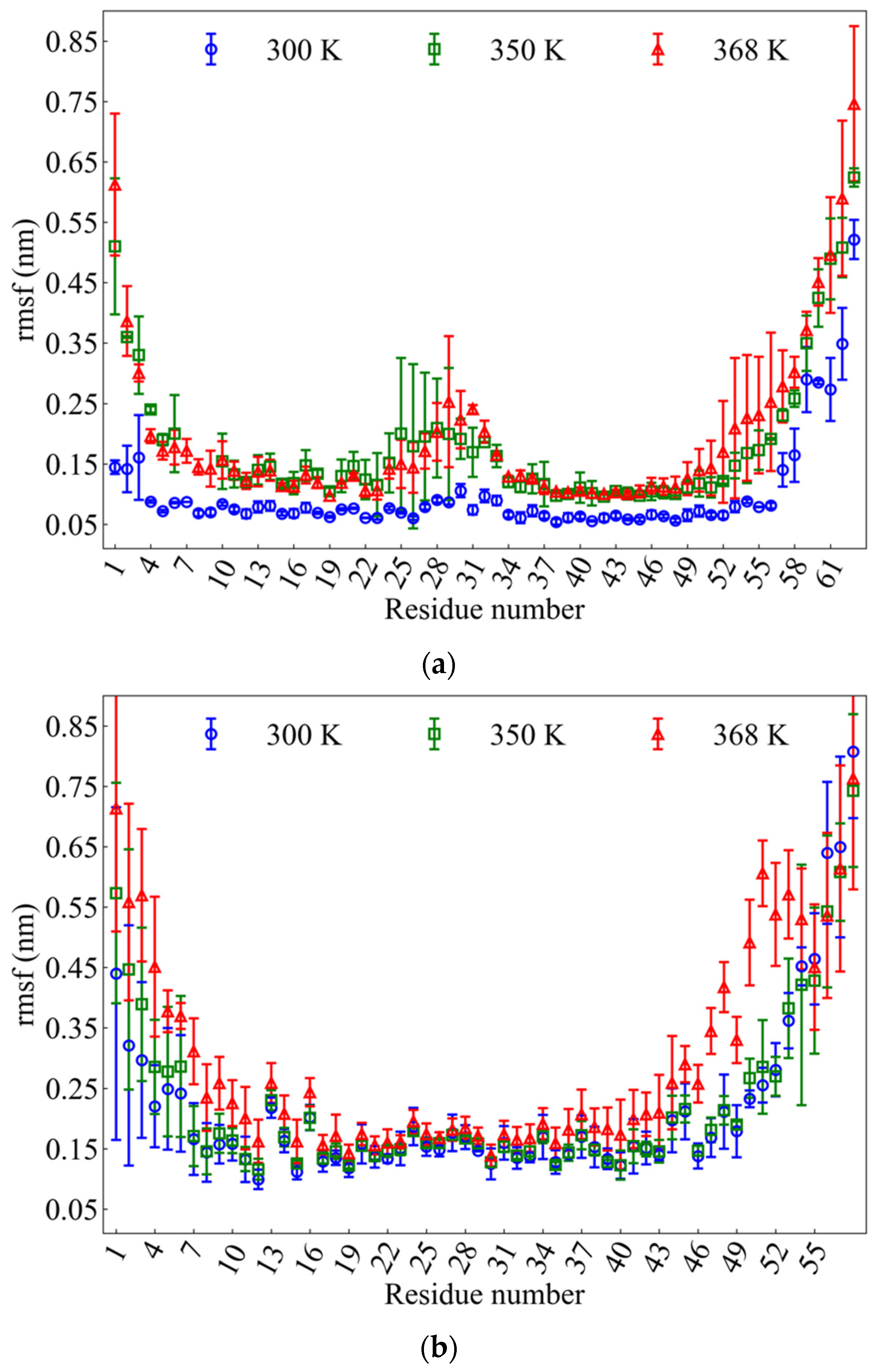
IJMS | Free Full-Text | Structure and Thermal Stability of wtRop and RM6 Proteins through All-Atom Molecular Dynamics Simulations and Experiments | HTML

Root Mean Square Fluctuation. Per-residue root mean square fluctuation... | Download Scientific Diagram

Calculating the root mean square fluctuation over a trajectory — MDAnalysis User Guide documentation
The role of disulfide bonds in a Solanum tuberosum saposin-like protein investigated using molecular dynamics | PLOS ONE

Fluctuations and Correlations in Crystalline Protein Dynamics: A Simulation Analysis of Staphylococcal Nuclease - ScienceDirect