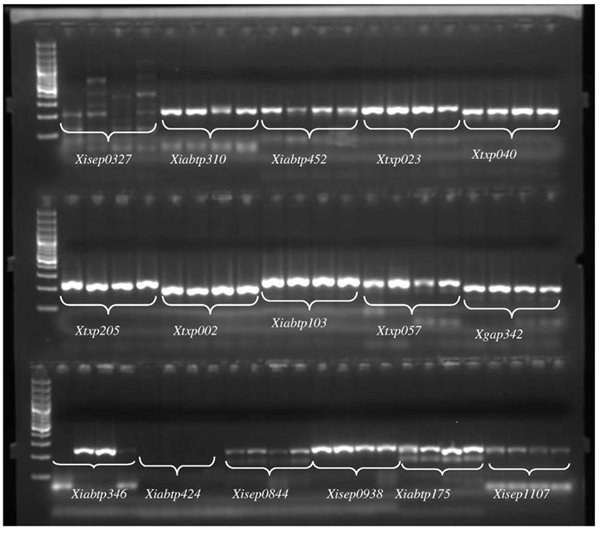
A general protocol for developing SSR markers with a SSR-enrichment step. | Download Scientific Diagram
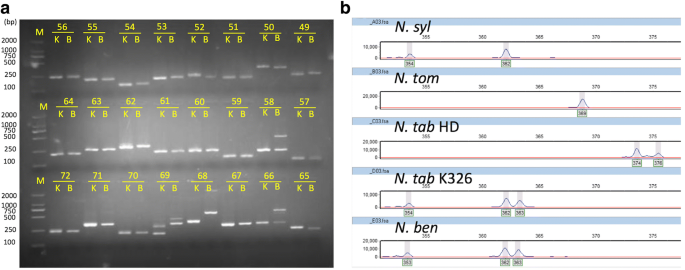
Comparative genome-wide characterization leading to simple sequence repeat marker development for Nicotiana | BMC Genomics | Full Text

SciELO - Brasil - Microsatellite markers: what they mean and why they are so useful Microsatellite markers: what they mean and why they are so useful

Development of unigene-derived SSR markers in cowpea (Vigna unguiculata) and their transferability to other Vigna species
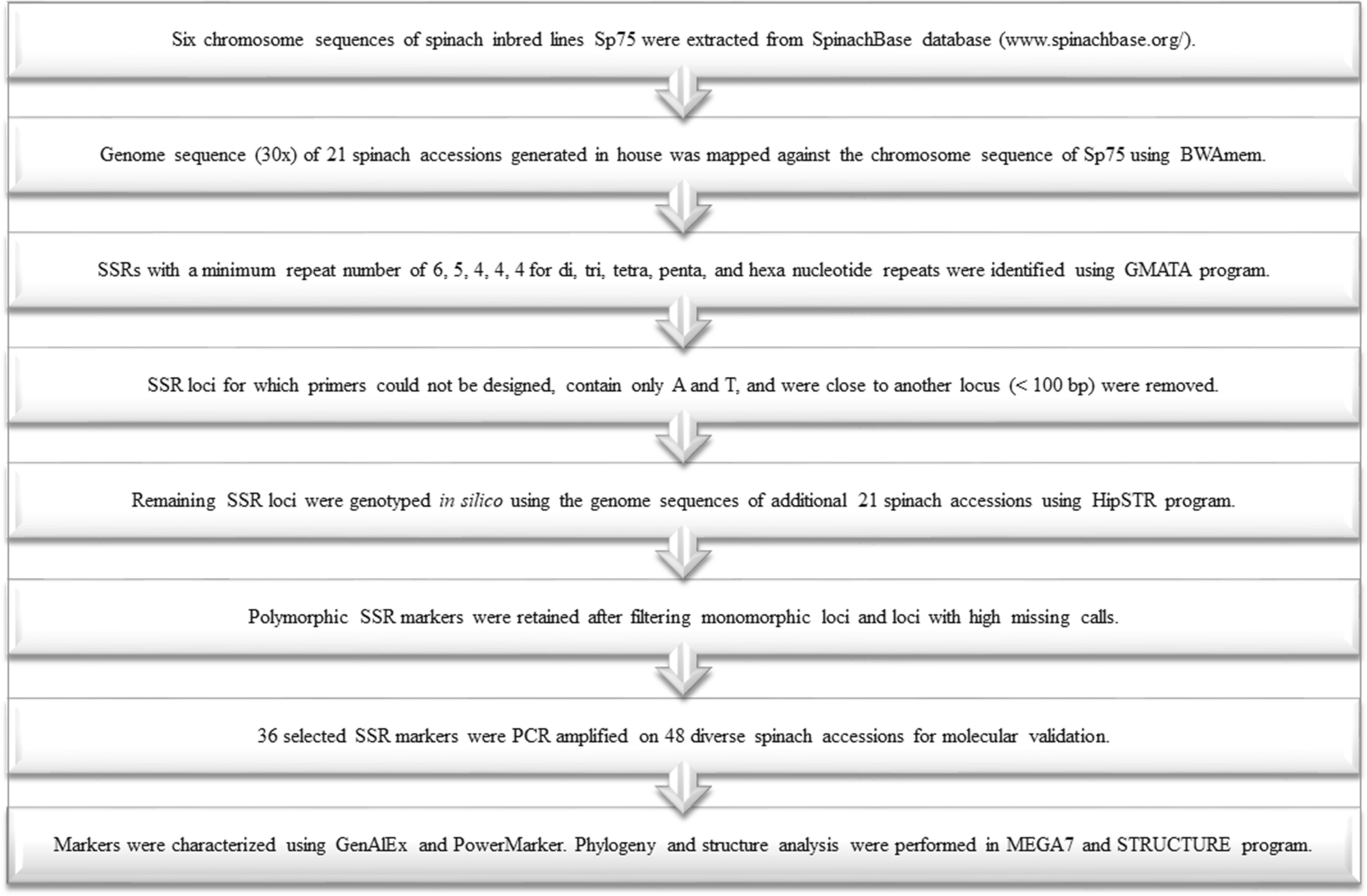
Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports

Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports

Development of EST–SSR markers in castor bean (Ricinus communis L.) and their utilization for genetic purity testing of hybrids
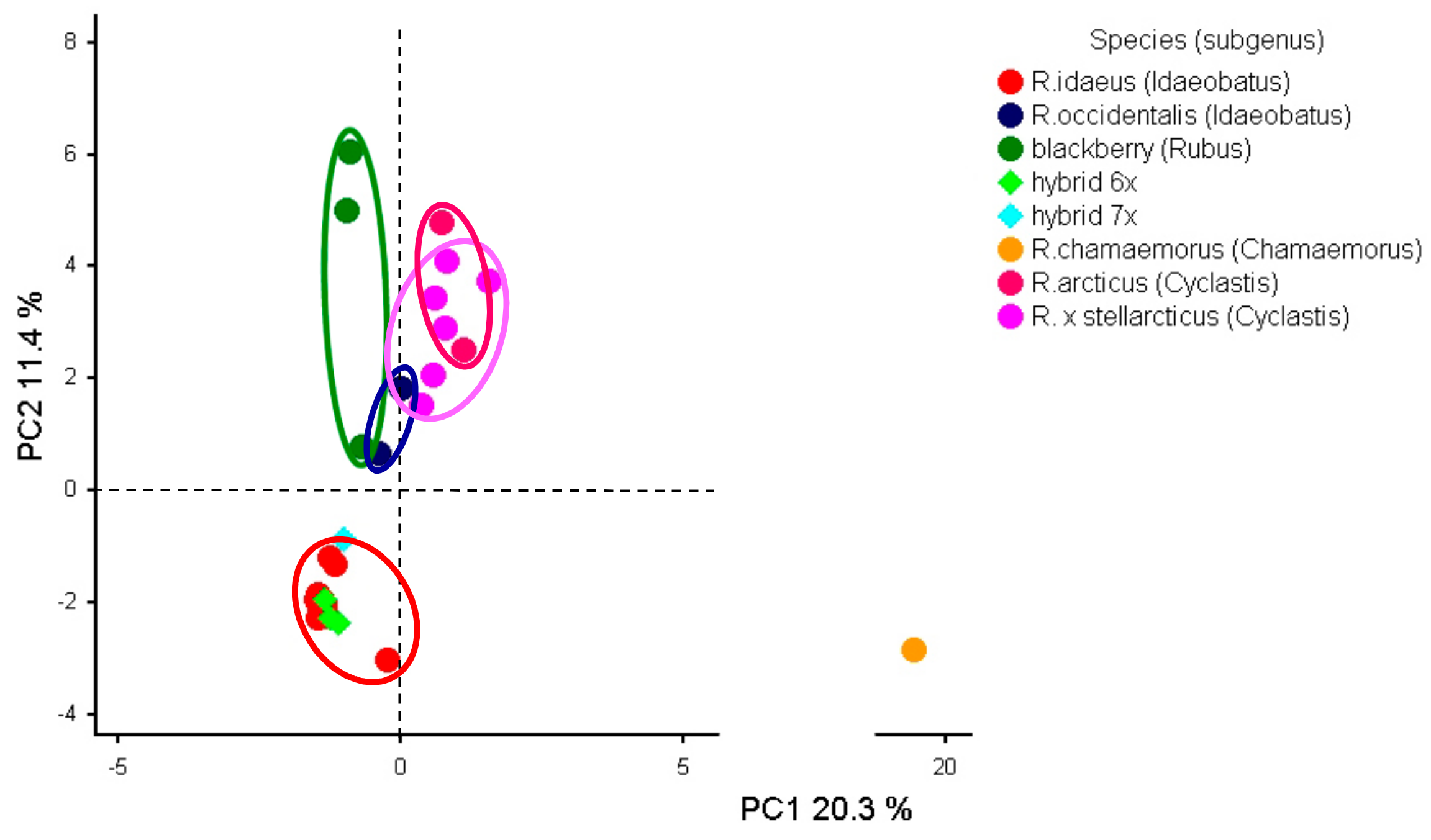
Genes | Free Full-Text | Transferability and Polymorphism of SSR Markers Located in Flavonoid Pathway Genes in Fragaria and Rubus Species
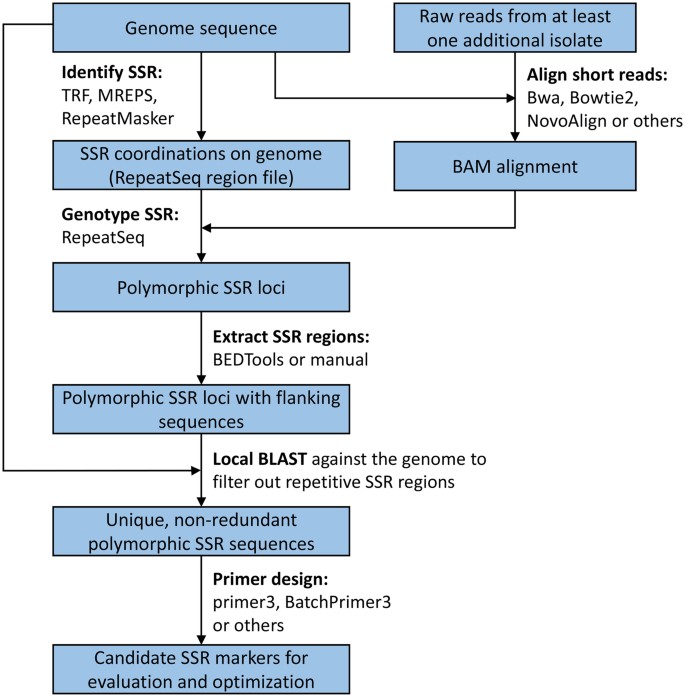
New microsatellite markers for population studies of Phytophthora cinnamomi, an important global pathogen | Scientific Reports
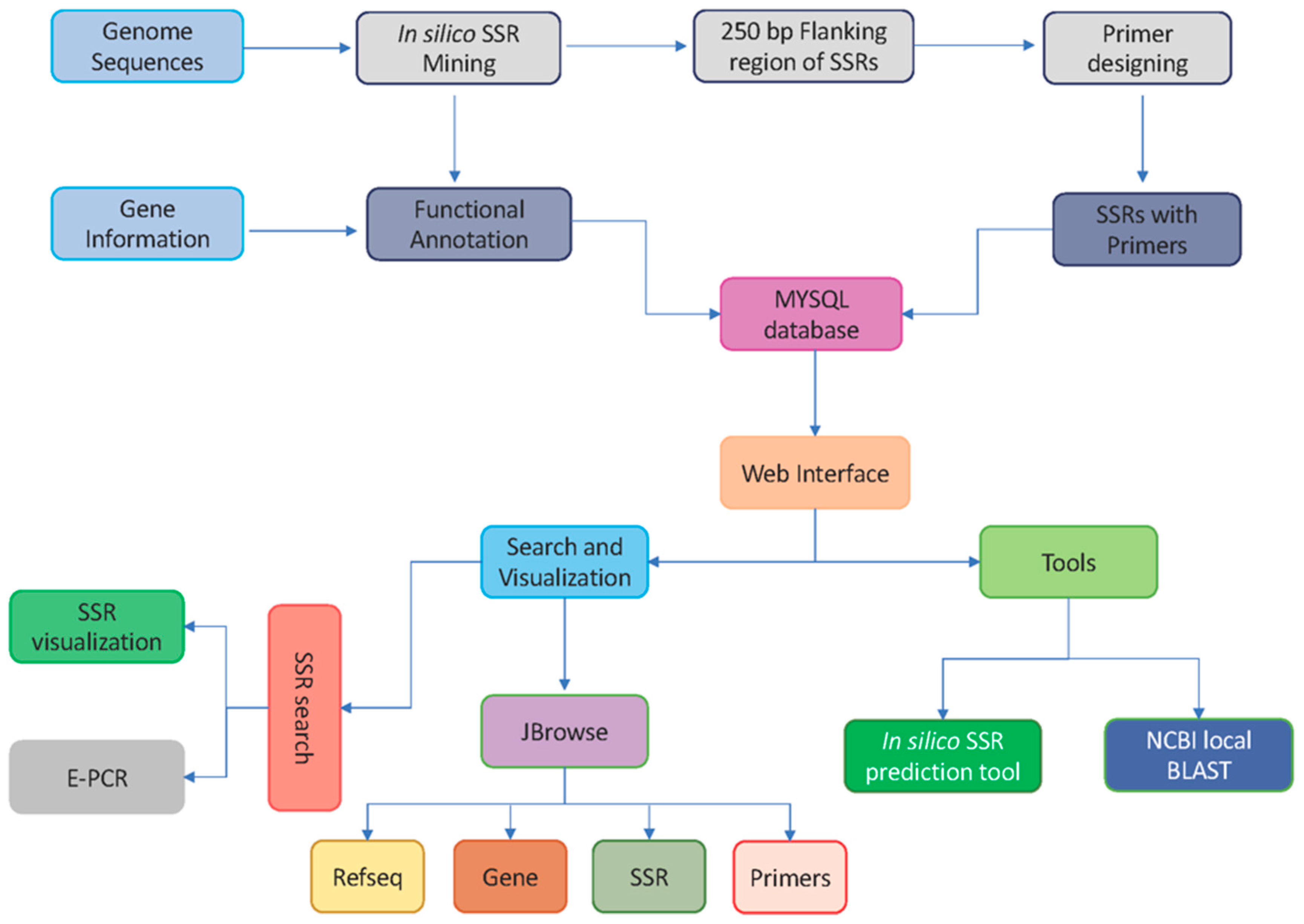
Genes | Free Full-Text | citSATdb: Genome-Wide Simple Sequence Repeat (SSR) Marker Database of Citrus Species for Germplasm Characterization and Crop Improvement | HTML
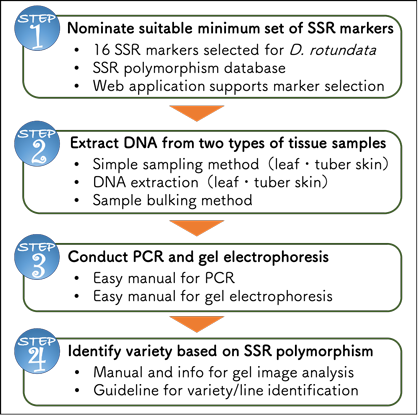
SSR marker technology package for variety identification of white Guinea Yam | Japan International Research Center for Agricultural Sciences | JIRCAS
Development of novel EST-SSR markers for ploidy identification based on de novo transcriptome assembly for Misgurnus anguillicaudatus | PLOS ONE
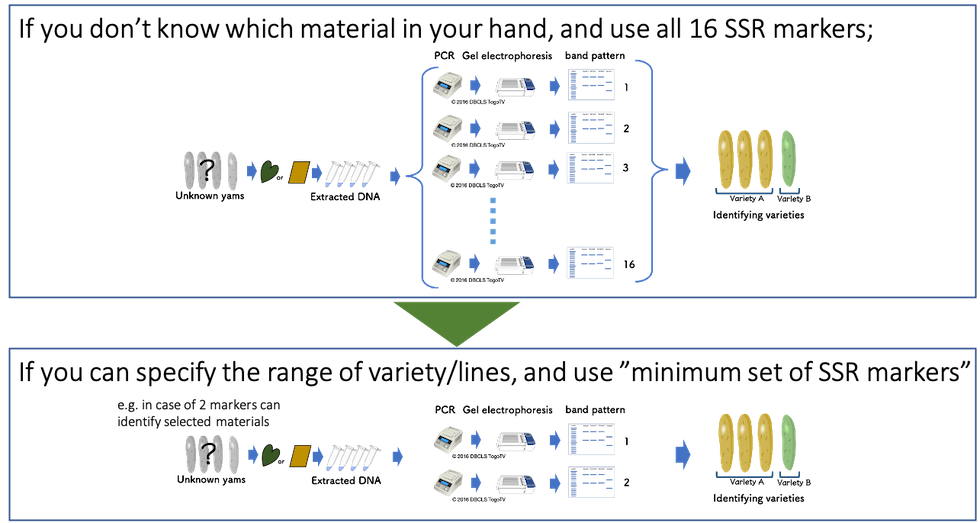
Yam Variety Identification Toolkit:Step 1 | Japan International Research Center for Agricultural Sciences | JIRCAS

Development of Genome-wide SSR Markers for Physical Map Construction with PCR-based Polymorphic SSRs in Jute (Corchorus Spp.) | SpringerLink

Identification and Sequence-Based Validation of the EST-SSR Markers from Calotropis procera | SpringerLink
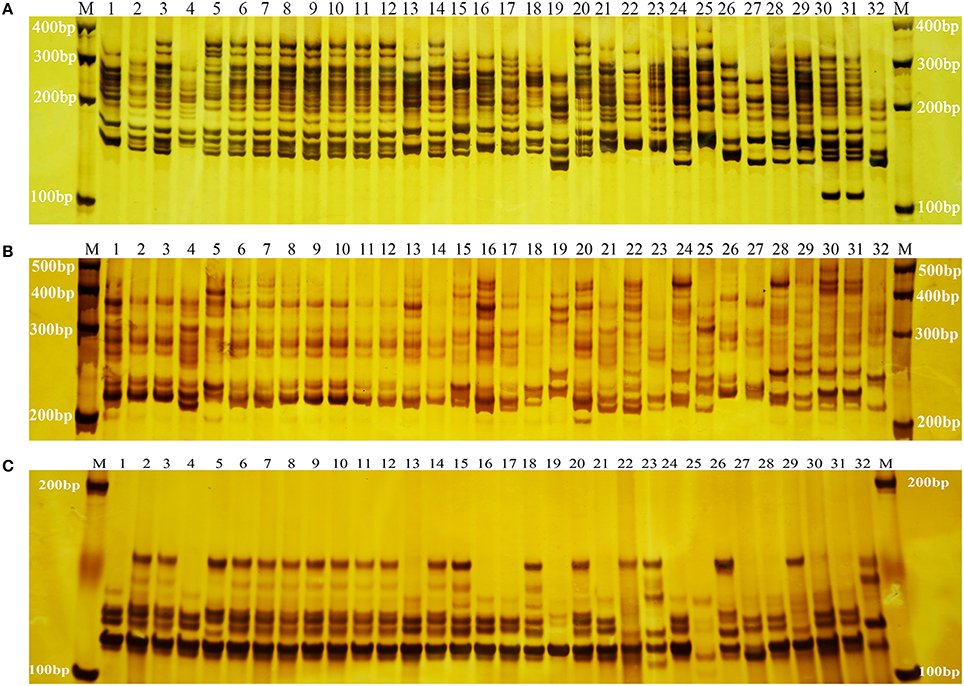
Frontiers | Development of SSR Markers and Assessment of Genetic Diversity in Medicinal Chrysanthemum morifolium Cultivars

EST-SSR markers derived from an elite barley cultivar (Hordeum vulgare L. 'Morex'): polymorphism and genetic marker potential. | Semantic Scholar