
Development and characterization of nitrogen and phosphorus use efficiency responsive genic and miRNA derived SSR markers in wheat | Heredity

Discovery of miRNAs and Development of Heat-Responsive miRNA-SSR Markers for Characterization of Wheat Germplasm for Terminal Heat Tolerance Breeding | Semantic Scholar
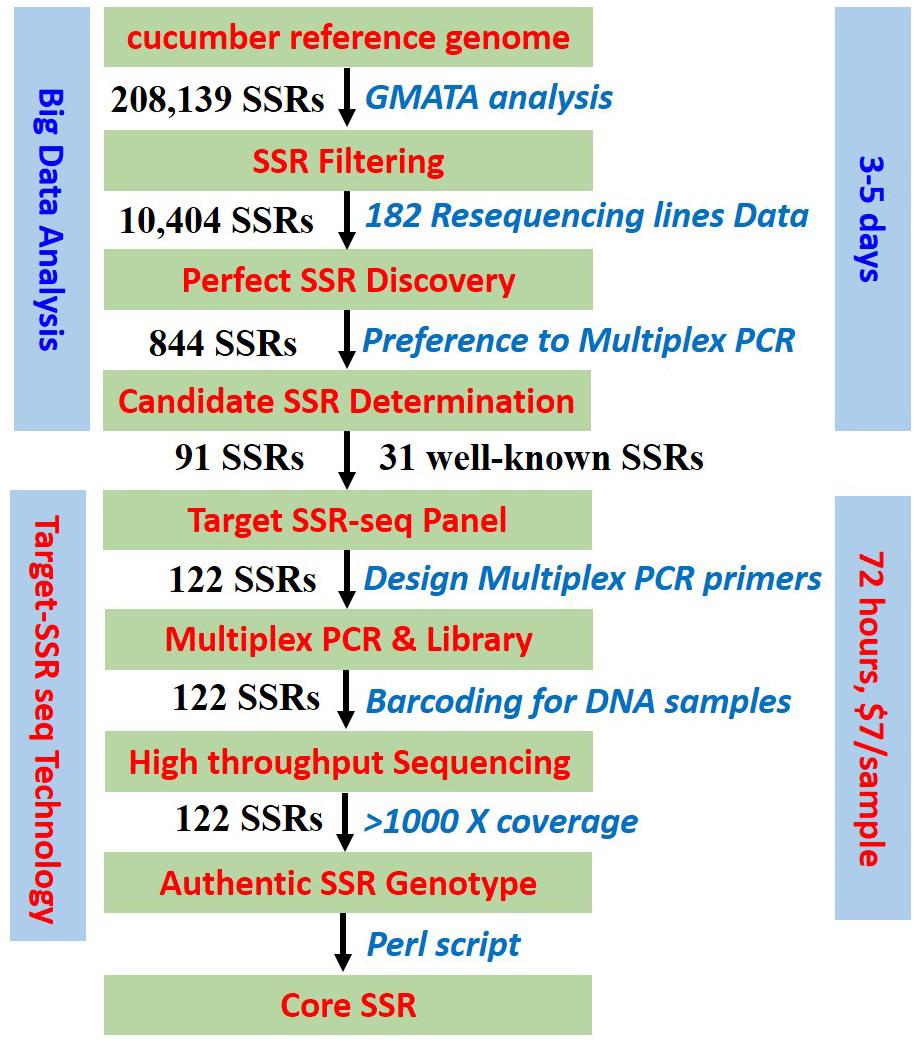
Frontiers | Target SSR-Seq: A Novel SSR Genotyping Technology Associate With Perfect SSRs in Genetic Analysis of Cucumber Varieties
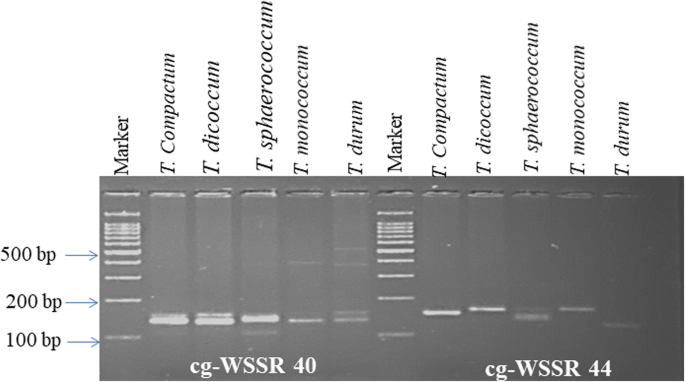
Identification, analysis and development of salt responsive candidate gene based SSR markers in wheat | BMC Plant Biology | Full Text
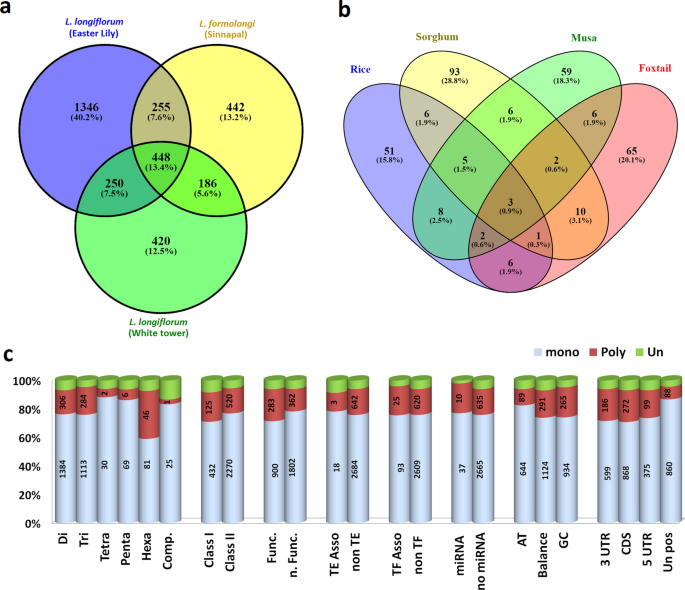
Transcriptome wide SSR discovery cross-taxa transferability and development of marker database for studying genetic diversity population structure of Lilium species | Scientific Reports
Genome-Wide Analysis of Microsatellite Markers Based on Sequenced Database in Chinese Spring Wheat (Triticum aestivum L.) | PLOS ONE

Linkage-physical maps of SSR markers and potential candidate genes. The... | Download Scientific Diagram

Development and Use of miRNA-derived SSR Markers to Study Genetic Diversity, Population Structure and Characterization of Genotypes for Heat Tolerance Breeding in Bread Wheat (Triticum aestivum L.) | bioRxiv
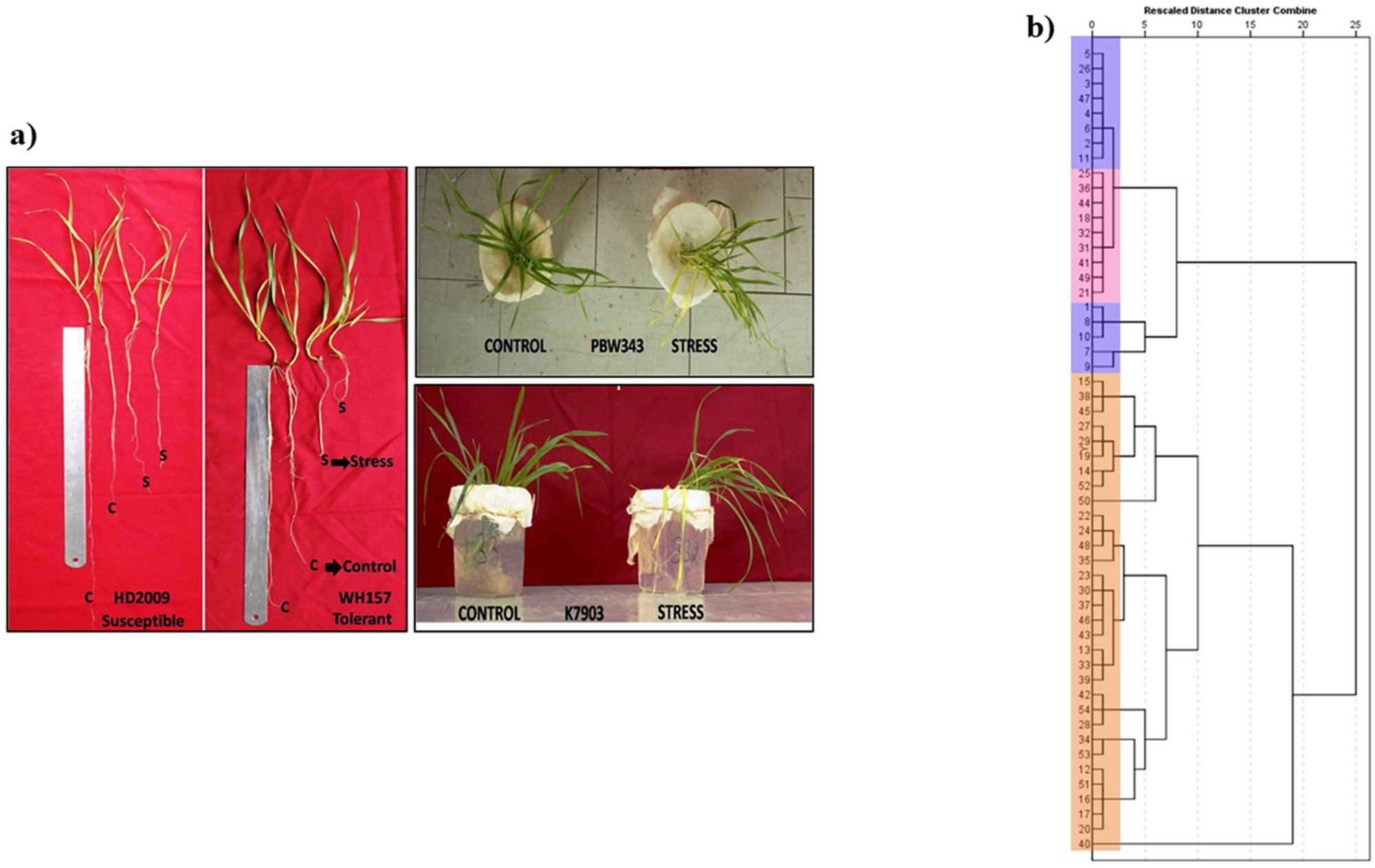
Identification and development of novel salt-responsive candidate gene based SSRs (cg-SSRs) and MIR gene based SSRs (mir-SSRs) in bread wheat (Triticum aestivum) | Scientific Reports

PDF) Development and mapping of microsatellite (SSR) markers in wheat | Sukhwinder Singh - Academia.edu

Map chart of 19 wheat chromosomes showing the associated SSR markers... | Download Scientific Diagram
![PDF] Interspecific transferability and comparative mapping of barley EST-SSR markers in wheat, rye and rice | Semantic Scholar PDF] Interspecific transferability and comparative mapping of barley EST-SSR markers in wheat, rye and rice | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/46df3d22fad48aafbb15d6bd9d2bb4401b3b92a4/3-Table1-1.png)
PDF] Interspecific transferability and comparative mapping of barley EST-SSR markers in wheat, rye and rice | Semantic Scholar

Development of RNA-seq-based molecular markers for characterizing Thinopyrum bessarabicum and Secale introgressions in wheat

EST SSR primers designed from contigs available in the coffee EST database. | Download Scientific Diagram
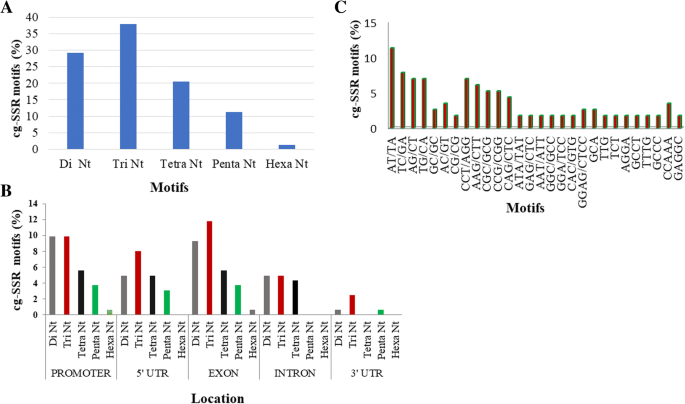
Identification, analysis and development of salt responsive candidate gene based SSR markers in wheat | BMC Plant Biology | Full Text
Genome-Wide Analysis of Microsatellite Markers Based on Sequenced Database in Chinese Spring Wheat (Triticum aestivum L.) | PLOS ONE
Genetic diversity, population structure and marker-trait associations for agronomic and grain traits in wild diploid wheat Triti
Development and use of miRNA-derived SSR markers for the study of genetic diversity, population structure, and characterization of genotypes for breeding heat tolerant wheat varieties | PLOS ONE
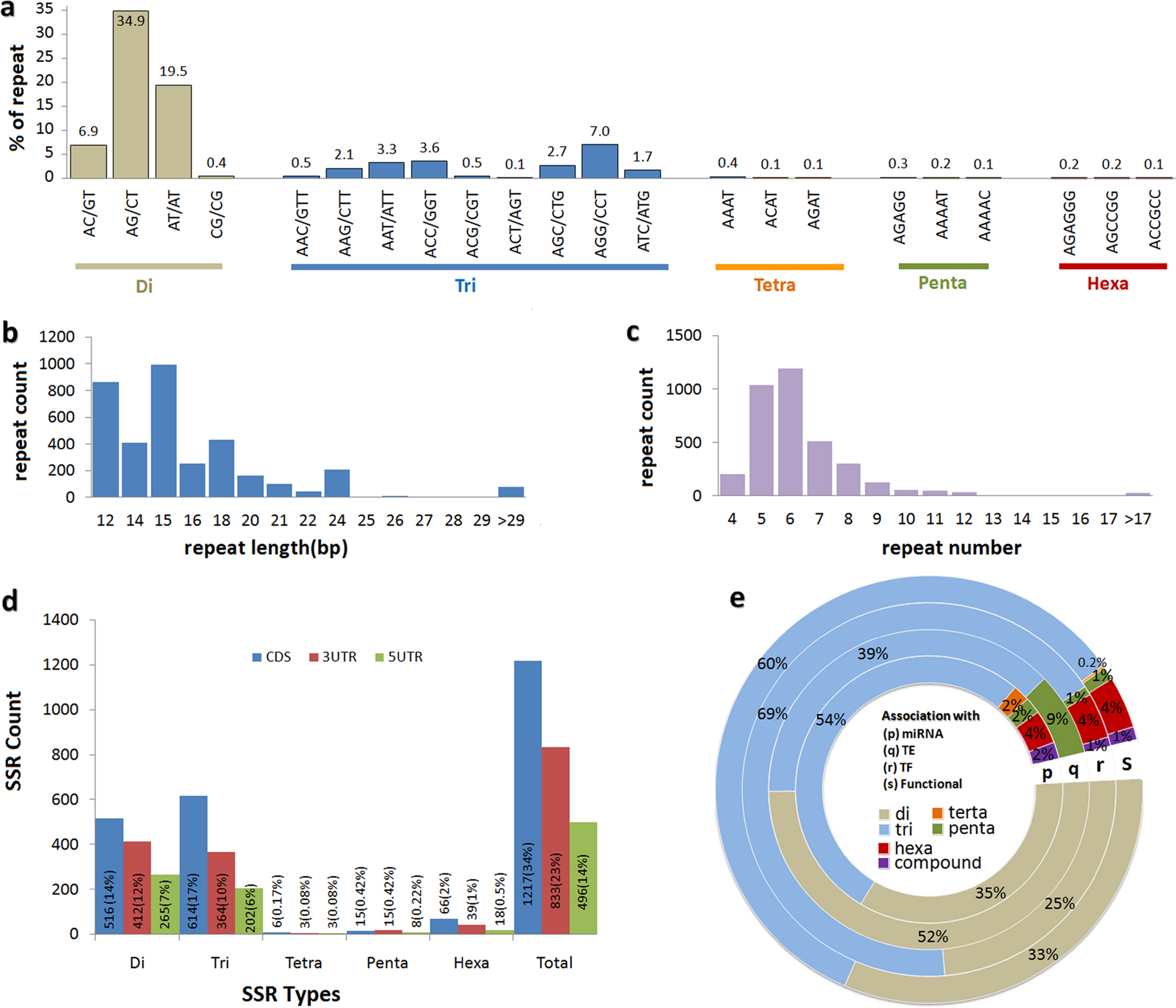
Transcriptome wide SSR discovery cross-taxa transferability and development of marker database for studying genetic diversity population structure of Lilium species | Scientific Reports

Comprehensive functional analysis and mapping of SSR markers in the chickpea genome (Cicer arietinum L.) - ScienceDirect
Genome-Wide Analysis of Microsatellite Markers Based on Sequenced Database in Chinese Spring Wheat (Triticum aestivum L.) | PLOS ONE
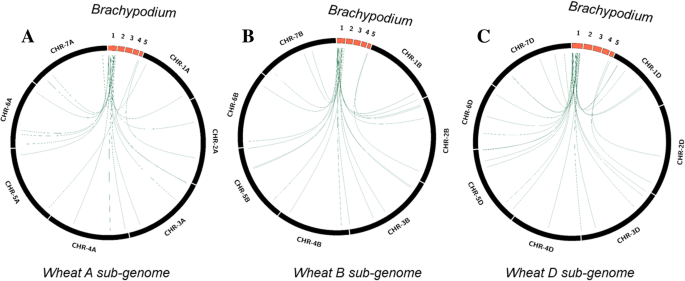
Identification, analysis and development of salt responsive candidate gene based SSR markers in wheat | BMC Plant Biology | Full Text
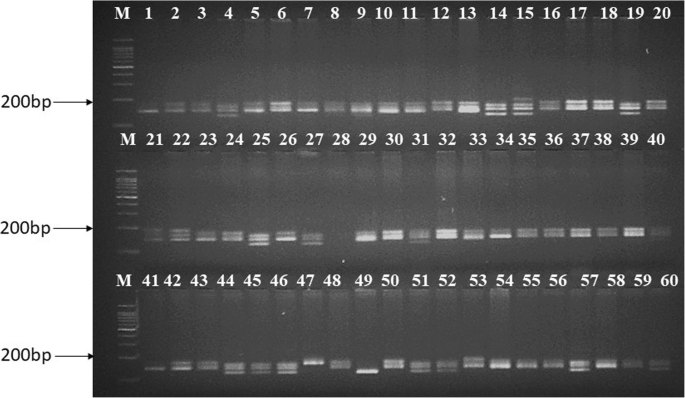
Identification, analysis and development of salt responsive candidate gene based SSR markers in wheat | BMC Plant Biology | Full Text

Frontiers | Putative Microsatellite DNA Marker-Based Wheat Genomic Resource for Varietal Improvement and Management
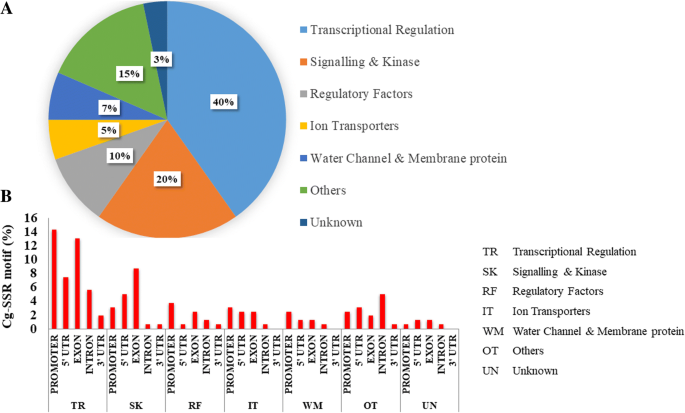
Identification, analysis and development of salt responsive candidate gene based SSR markers in wheat | BMC Plant Biology | Full Text


